Overview: De novo antibody sequencing is to determine the amino acid sequence of an antibody without prior knowledge of its genetic sequence or access to its cDNA or original cell line. You will need the powerful tool when B cells or any sequence information are not available. The mass spectrometry-based de novo antibody sequencing methods are less costly and time-consuming than traditional sequencing methods, such as Edman degradation.
Sample preparation: The antibody sample is usually purified and then denatured, reduced, and alkylated to break and blocked disulfide bonds. The antibody sample is then digested into overlapping peptides using multiple enzymes such as Pepsin, Trypsin, Chymotrypsin, Asp N, Lys C and so on.
High-performance liquid chromatography – mass spectrometry (HPLC-MS) analysis: The resulting peptide mixtures are separated using reversed phase (RP) high-performance liquid chromatography (HPLC). The peptides are ionized and analyzed using mass spectrometry to determine the mass-to-charge ratio of each parental and fragmental ions, where both HCD and ETD are applied.
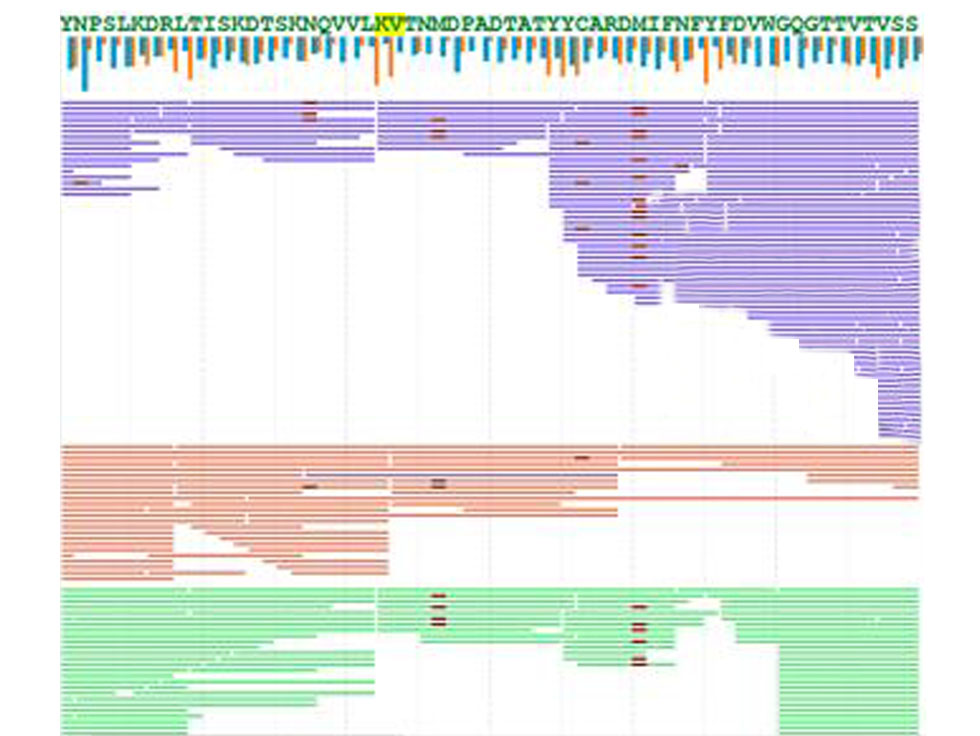
Data analysis: The mass spectra data are analyzed using de novo sequencing algorithms to acquire the sequence of each peptide, where Leucine (L) and Isoleucine (I) are differentiated by a couple of methods, such as enzyme preference and unique w ions. Then the overlapping peptide sequences are assembled into a complete antibody sequence.
Verification: The determined antibody sequence is then verified by the intact mass results.